Rapid Microbiological Methods (RMMs) are technologies developed to detect, enumerate, or identify microorganisms faster than conventional culture-based techniques.
Traditional microbiological tests rely on visible colony formation, which may take several days to grow. In contrast, RMMs reduce detection time by measuring microbial growth, viability, cellular components, or genetic material using advanced analytical tools.
Regulatory authorities such as the European Pharmacopoeia (Ph. Eur. 5.1.6) and USP <1223> recognize multiple technological approaches for rapid microbiological testing. Although classifications may vary slightly, RMMs are generally grouped into the following categories:
- Growth-based methods
- Direct measurement methods
- Cell component analysis
- Optical spectroscopy methods
- Nucleic acid amplification technologies
- Microelectromechanical systems (MEMS) and microarray platforms
Let us go through each category in detail.
1. Growth-Based Rapid Methods
Growth-based RMMs depend on microbial replication, but they detect growth much earlier by measuring biochemical changes before visible colonies appear.
These methods continue to use conventional liquid or solid media.
Impedance Microbiology
Impedance systems detect changes in the electrical properties of culture media during microbial growth. As microorganisms metabolize nutrients, they produce ionic by-products that alter the conductivity of the medium.
An increase in conductance results in a measurable decrease in impedance. These electrical changes correlate with microbial growth and allow earlier detection compared to traditional visual methods.
Carbon Dioxide Detection Systems
Actively growing microorganisms produce carbon dioxide (CO₂) as a metabolic by-product.
In colorimetric CO₂ detection systems, a sensor is placed within the culture bottle but separated from the medium by a semipermeable membrane. CO₂ diffuses into the sensor, dissolves in water, and generates hydrogen ions. This leads to a measurable color change that is detected photometrically.
Biochemical and Substrate Utilization Systems
Some automated systems detect microbial growth based on the utilization of specific carbohydrates or enzymatic substrates incorporated into the media. Metabolic reactions generate measurable colorimetric or fluorometric signals.
Rapid Microcolony Detection (Digital Imaging)
In conventional plate methods, visible colonies may require several days to form. Rapid microcolony detection systems use laser scanning and digital imaging to detect very small colonies on membrane filters at an earlier stage.
These systems allow enumeration within a shorter time frame while maintaining organism viability for subsequent identification.
Fluorescent Staining of Microcolonies
Microcolonies growing on solid media can be stained with fluorescent dyes and detected using laser excitation. This approach enables earlier visualization compared to manual inspection.
Growth-based RMMs still depend on microbial replication. They reduce detection time but do not eliminate the need for growth.
2. Direct Measurement Methods
Direct measurement technologies detect and differentiate individual microbial cells without requiring prior growth.
These methods use viability stains and laser-based detection systems.
Flow Cytometry
Flow cytometry counts individual particles as they pass through a laser beam. Microorganisms are stained using fluorescent dyes such as: Propidium iodide (penetrates cells with damaged membranes) Thiazole orange (penetrates all cells)
By combining dyes, the system can differentiate between cells with intact membranes and membrane-compromised cells.
However, it is important to understand that membrane integrity does not always correlate perfectly with replicative ability. Therefore, the method must be validated to establish suitability for the intended purpose.
Solid-Phase Cytometry
In this method, microorganisms are captured on a membrane, stained, and detected by laser scanning. Individual fluorescent events are counted.
3. Cell Component Analysis
These methods detect specific cellular components as indirect indicators of microbial presence. They do not necessarily confirm microbial growth.
ATP Bioluminescence
Adenosine triphosphate (ATP) is present in all living cells. In ATP bioluminescence assays, ATP reacts with luciferin in the presence of the enzyme luciferase.
This reaction produces light (bioluminescence), not fluorescence. The emitted photons are measured using a luminometer.
ATP-based systems are widely used for hygiene monitoring and rapid contamination assessment. However, they do not distinguish microbial ATP from other biological ATP unless additional sample preparation steps are used.
Endotoxin Testing (LAL)
Limulus Amebocyte Lysate (LAL) testing detects bacterial endotoxin (lipopolysaccharide, LPS) from Gram-negative bacteria.
It does not detect viable microorganisms.
Therefore, the LAL test is a rapid pyrogen test, not a microbial detection method. It indicates the presence of endotoxin but does not confirm active contamination.
Fatty Acid Analysis
Microorganisms possess characteristic fatty acid profiles. Analytical techniques such as gas chromatography can generate fingerprint patterns used for microbial identification.
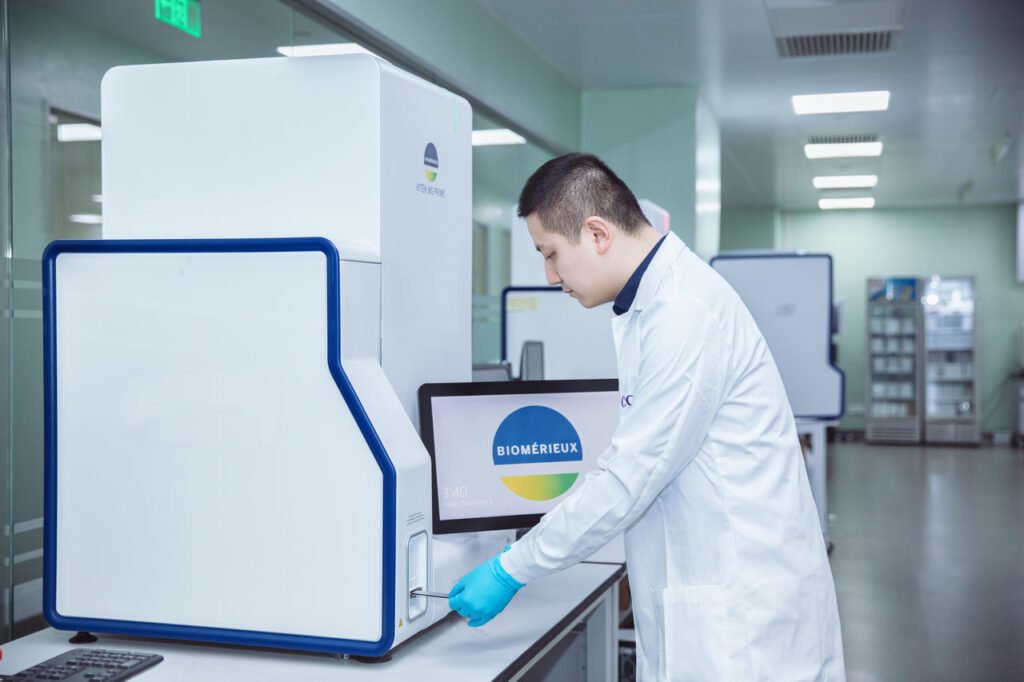
MALDI-TOF Mass Spectrometry
Matrix-Assisted Laser Desorption Ionization–Time of Flight (MALDI-TOF) mass spectrometry is used for rapid microbial identification.
It analyzes protein spectral fingerprints of microorganisms. However, it requires isolated colonies grown on solid media before analysis. Therefore, it is a rapid identification tool rather than a primary detection method.

4. Optical Spectroscopy Methods
Optical systems use light scattering and fluorescence principles to detect particles and biological signals.
Real-Time Biofluorescent Air Monitoring
These systems draw air into an instrument where particles pass through a laser beam. Particle size is determined using Mie scattering principles.
Particles containing biological molecules such as NADH, riboflavin, or dipicolinic acid may emit autofluorescence signals when excited by specific wavelengths.
These systems detect biological particles in real time. However, they do not confirm viability and may detect non-viable biological fragments or environmental materials such as pollen.
Therefore, they are primarily used for environmental monitoring in cleanrooms.
5. Nucleic Acid Amplification Technologies
These methods detect microbial genetic material and do not require cellular growth.
Polymerase Chain Reaction (PCR)
PCR amplifies specific DNA sequences, producing millions of copies within a short period. It allows highly sensitive and specific detection of targeted microorganisms.
PCR detects DNA that originates from viable or non-viable organisms.
Reverse Transcriptase PCR (RT-PCR)
RT-PCR targets RNA, often messenger RNA (mRNA), which may correlate more closely with viable organisms due to its shorter half-life.
Riboprinting
Riboprinting is an automated method based on restriction fragment length polymorphism (RFLP) analysis of rRNA genes. The resulting patterns are compared with databases for identification.
It is distinct from direct 16S sequencing.
Gene Sequencing
Gene sequencing determines the exact nucleotide order of amplified DNA fragments. PCR is used for amplification prior to gene sequencing.
Sequencing gives highly accurate identification at the genus and species levels.
6. Microelectromechanical Systems (MEMS) and Microarrays
MEMS technologies integrate microfluidics, biosensors, and miniaturized analytical systems.
DNA Microarrays
Microarrays are based on nucleic acid hybridization principles. Extracted DNA is amplified and fluorescently labeled, then hybridized to probes immobilized on a chip.
The chip contains species-specific or gene-specific targets. Hybridization patterns enable identification of microorganisms, including applications such as mycoplasma detection.
Microarrays represent a “lab-on-a-chip” approach combining amplification, detection, and analysis in a miniaturized platform.
RMMs do not automatically replace traditional microbiological methods. Their suitability depends on the intended application, product type, and regulatory acceptance.
When properly validated and implemented, rapid microbiological methods represent a powerful advancement in pharmaceutical microbiology.
Selecting Surfaces for Disinfectant Efficacy Testing
Fluid Thioglycollate Medium (FTM) in Sterility Testing
Soybean Casein Digest Medium (SCDM) and Its Role in Media Fills



Do regulators allow RMMs for finished product testing?
Yes. Regulators do allow Rapid Microbiological Methods for finished product testing, provided they are properly validated and scientifically justified.